Large-scale discovery of chromatin dysregulation induced by oncofusions and other protein-coding variants
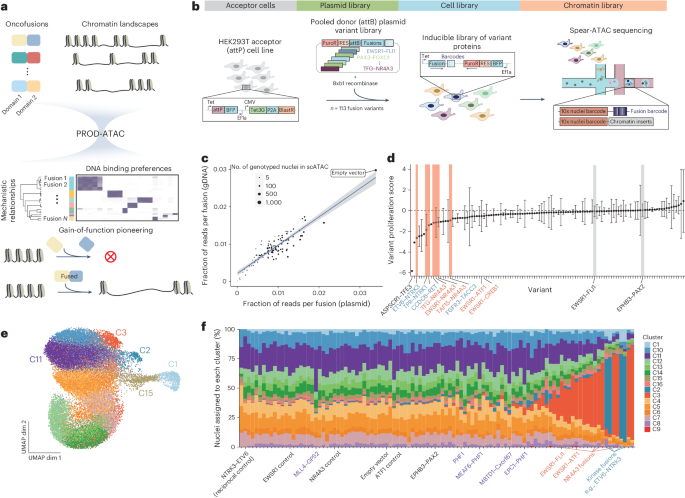
Technology tamfitronics
Przybyla, L. & Gilbert, L. A. A new era in functional genomics screens. Night. Rev. The gene. 2389–103 (2021).
Findlay, G. M. Linking genome variants to disease: scalable approaches to test the functional impact of human mutations. Hum. Mol. Genet. 30R187–R197 (2021).
Findlay, G. M. et al. Accurate classification of BRCA1 variants with saturation genome editing. Nature 562217–222 (2018).
Tewhey, R. et al. Direct identification of hundreds of expression-modulating variants using a multiplexed reporter assay. Cell 1651519–1529 (2016).
van Arensbergen, J. et al. High-throughput identification of human SNPs affecting regulatory element activity. Nat. Genet. 511160–1169 (2019).
Doench, J. G. Am I ready for CRISPR? A user’s guide to genetic screens. Night. Rev. The gene. 1967–80 (2018).
Fowler, D. M. & Fields, S. Deep mutational scanning: a new style of protein science. Nat. Methods 11801–807 (2014).
Replogle, J. M. et al. Combinatorial single-cell CRISPR screens by direct guide RNA capture and targeted sequencing. Nat. Biotechnol. 38954–961 (2020).
Adamson, B. et al. A multiplexed single-cell CRISPR screening platform enables systematic dissection of the unfolded protein response. Cell 1671867–1882 (2016).
Dixit, A. et al. Perturb-seq: dissecting molecular circuits with scalable single-cell RNA profiling of pooled genetic screens. Cell 1671853–1866 (2016).
Datlinger, P. et al. Pooled CRISPR screening with single-cell transcriptome readout. Nat. Methods 14297–301 (2017).
Kim, H. S. et al. Direct measurement of engineered cancer mutations and their transcriptional phenotypes in single cells. Nat. Biotechnol. https://doi.org/10.1038/s41587-023-01949-8 (2023).
Ursu, O. et al. Massively parallel phenotyping of coding variants in cancer with Perturb-seq. Nat. Biotechnol. 40896–905 (2022).
Pierce, S. E., Granja, J. M. & Greenleaf, W. J. High-throughput single-cell chromatin accessibility CRISPR screens enable unbiased identification of regulatory networks in cancer. Nat. Commun. 122969 (2021).
Mertens, F., Johansson, B., Fioretos, T. & Mitelman, F. The emerging complexity of gene fusions in cancer. Night. Rev. Cancer 15371–381 (2015).
Mertens, F., Antonescu, C. R. & Mitelman, F. Gene fusions in soft tissue tumors: recurrent and overlapping pathogenetic themes. Genes Chromosomes Cancer 55291–310 (2016).
Gryder, B. E. et al. PAX3–FOXO1 establishes myogenic super enhancers and confers BET bromodomain vulnerability. Cancer Discov. 7884–899 (2017).
Riggi, N. et al. EWS-FLI1 utilizes divergent chromatin remodeling mechanisms to directly activate or repress enhancer elements in Ewing sarcoma. Cancer Cell 26668–681 (2014).
Boulay, G. et al. Cancer-specific retargeting of BAF complexes by a prion-like domain. Cell 171163–178 (2017).
Jang, Y. E. et al. ChimerDB 4.0: an updated and expanded database of fusion genes. Nucleic Acids Res. 48D817–D824 (2020).
Sweeney, N. P. & Vink, C. A. The impact of lentiviral vector genome size and producer cell genomic to gag-pol mRNA ratios on packaging efficiency and titre. Mol. Ther. Methods Clin. Dev. 21574–584 (2021).
Milone, M. C. & O’Doherty, U. Clinical use of lentiviral vectors. Leukemia 321529–1541 (2018).
Xie, S., Cooley, A., Armendariz, D., Zhou, P. & Hon, G. C. Frequent sgRNA-barcode recombination in single-cell perturbation assays. PLoS ONE 13e0198635 (2018).
Adamson, B., Norman, T. M., Jost, M. & Weissman, J. S. Approaches to maximize sgRNA-barcode coupling in Perturb-seq screens. Preprint at bioRxiv https://doi.org/10.1101/298349 (2018).
Feldman, D., Singh, A., Garrity, A. J. & Blainey, P. C. Lentiviral co-packaging mitigates the effects of intermolecular recombination and multiple integrations in pooled genetic screens. Preprint at bioRxiv https://doi.org/10.1101/262121 (2018).
Parekh, U. et al. Mapping cellular reprogramming via pooled overexpression screens with paired fitness and single-cell RNA-sequencing readout. Cell Syst. 7548–555 (2018).
Hill, A. J. et al. On the design of CRISPR-based single-cell molecular screens. Nat. Methods 15271–274 (2018).
Matreyek, K. A., Stephany, J. J., Chiasson, M. A., Hasle, N. & Fowler, D. M. An improved platform for functional assessment of large protein libraries in mammalian cells. Nucleic Acids Res. 48e1 (2020).
Sánchez-Molina, S. et al. RING1B recruits EWSR1-FLI1 and cooperates in the remodeling of chromatin necessary for Ewing sarcoma tumorigenesis. Sci. Adv. 6eaba3058 (2020).
Deng, Q. et al. Oncofusion-driven de novo enhancer assembly promotes malignancy in Ewing sarcoma via aberrant expression of the stereociliary protein LOXHD1. Cell Rep. 39110971 (2022).
Manceau, L. et al. Divergent transcriptional and transforming properties of PAX3-FOXO1 and PAX7-FOXO1 paralogs. PLoS Genet. 18e1009782 (2022).
Orth, M. F. et al. Systematic multi-omics cell line profiling uncovers principles of Ewing sarcoma fusion oncogene-mediated gene regulation. Cell Rep. 41111761 (2022).
Tate, J. G. et al. COSMIC: the Catalogue Of Somatic Mutations In Cancer. Nucleic Acids Res. 47D941–D947 (2019).
Granja, J. M. et al. ArchR is a scalable software package for integrative single-cell chromatin accessibility analysis. Nat. Genet. 53403–411 (2021).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 1843573–3587 (2021).
Jost, M. et al. Titrating gene expression using libraries of systematically attenuated CRISPR guide RNAs. Nat. Biotechnol. 38355–364 (2020).
Zhang, Y. et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. 9R137 (2008).
Sunkel, B. D. et al. Evidence of pioneer factor activity of an oncogenic fusion transcription factor. iScience 24102867 (2021).
AACR Project GENIE Consortium AACR Project GENIE: powering precision medicine through an international consortium. Cancer Discov. 7818–831 (2017).
Chang, W.-I. et al. Molecular targets for novel therapeutics in pediatric fusion-positive non-CNS solid tumors. Front. Pharmacol. 12747895 (2022).
Perry, J. A., Seong, B. K. A. & Stegmaier, K. Biology and therapy of dominant fusion oncoproteins involving transcription factor and chromatin regulators in sarcomas. Annu. Rev. Cancer Biol. 3299–321 (2019).
Möller, E. et al. EWSR1-ATF1 dependent 3D connectivity regulates oncogenic and differentiation programs in clear cell sarcoma. Nat. Commun. 132267 (2022).
Johnson, K. M. et al. Role for the EWS domain of EWS/FLI in binding GGAA-microsatellites required for Ewing sarcoma anchorage independent growth. Proc. Natl Acad. Sci. USA 1149870–9875 (2017).
Johnson, K. M., Taslim, C., Saund, R. S. & Lessnick, S. L. Identification of two types of GGAA-microsatellites and their roles in EWS/FLI binding and gene regulation in Ewing sarcoma. PLoS ONE 12e0186275 (2017).
Guillon, N. et al. The oncogenic EWS-FLI1 protein binds in vivo GGAA microsatellite sequences with potential transcriptional activation function. PLoS ONE 4e4932 (2009).
Li, Z. et al. ETV6–NTRK3 fusion oncogene initiates breast cancer from committed mammary progenitors via activation of AP1 complex. Cancer Cell 12542–558 (2007).
Przybyl, J. et al. Gene expression profiling of low-grade endometrial stromal sarcoma indicates fusion protein-mediated activation of the Wnt signaling pathway. Gynecol. Oncol. 149388–393 (2018).
Gordon, A. T. et al. A novel and consistent amplicon at 13q31 associated with alveolar rhabdomyosarcoma. Genes Chromosomes Cancer 28220–226 (2000).
Yoon, J. W., Lamm, M., Chandler, C., Iannaccone, P. & Walterhouse, D. Up-regulation of GLI1 in vincristine-resistant rhabdomyosarcoma and Ewing sarcoma. BMC Cancer 20511 (2020).
Birdsey, G. M. et al. The endothelial transcription factor ERG promotes vascular stability and growth through Wnt/β-catenin signaling. Dev. Cell 3282–96 (2015).
Brcic, I. et al. Implementation of copy number variations-based diagnostics in morphologically challenging EWSR1/FUS::NFATC2 neoplasms of the bone and soft tissue. Int. J. Mol. Sci. 2316196 (2022).
Deplus, R. et al. TMPRSS2-ERG fusion promotes prostate cancer metastases in bone. Oncotarget 811827–11840 (2016).
Parviz, F. et al. Hepatocyte nuclear factor 4α controls the development of a hepatic epithelium and liver morphogenesis. Nat. Genet. 34292–296 (2003).
Zhang, B. et al. Proteogenomic characterization of human colon and rectal cancer. Nature 513382–387 (2014).
Weinstein, J. N. et al. The Cancer Genome Atlas Pan-Cancer analysis project. Nat. Genet. 451113–1120 (2013).
Davis, R. B., Kaur, T., Moosa, M. M. & Banerjee, P. R. FUS oncofusion protein condensates recruit mSWI/SNF chromatin remodeler via heterotypic interactions between prion‐like domains. Protein Sci. Publ. Protein Soc. 301454–1466 (2021).
Domingo, J. et al. Non-linear transcriptional responses to gradual modulation of transcription factor dosage. Preprint at bioRxiv https://doi.org/10.1101/2024.03.01.582837 (2024).
Backman, J. D. et al. Exome sequencing and analysis of 454,787 UK Biobank participants. Nature 599628–634 (2021).
Lancaster, A. K., Nutter-Upham, A., Lindquist, S. & King, O. D. PLAAC: a web and command-line application to identify proteins with prion-like amino acid composition. Bioinformatics 302501–2502 (2014).
Farag, M., Borcherds, W. M., Bremer, A., Mittag, T. & Pappu, R. V. Phase separation in mixtures of prion-like low complexity domains is driven by the interplay of homotypic and heterotypic interactions. Preprint at bioRxiv https://doi.org/10.1101/2023.03.15.532828 (2023).
Boncella, A. E. et al. Composition-based prediction and rational manipulation of prion-like domain recruitment to stress granules. Proc. Natl Acad. Sci. USA 1175826–5835 (2020).
Sprunger, M. L. & Jackrel, M. E. Prion-like proteins in phase separation and their link to disease. Biomolecules 111014 (2021).
Wang, Y. et al. Dissolution of oncofusion transcription factor condensates for cancer therapy. Nat. Chem. Biol. 191223–1234 (2023).
Frankish, A. et al. GENCODE 2021. Nucleic Acids Res. 49D916–D923 (2021).
Rubin, A. F. et al. A statistical framework for analyzing deep mutational scanning data. Genome Biol. 18150 (2017).
Corces, M. R. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14959–962 (2017).
Reske, J. J., Wilson, M. R. & Chandler, R. L. ATAC-seq normalization method can significantly affect differential accessibility analysis and interpretation. Epigenetics Chromatin 1322 (2020).
Center for High Throughput Computing (University of Wisconsin–Madison); https://doi.org/10.21231/GNT1-HW21. Accessed September 2021
Frenkel, M., Corban, J. E., Hujoel, M. L. A., Morris, Z. & Raman, S. Large-scale discovery of chromatin dysregulation induced by oncofusions and other protein-coding variants. NCBI https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE243553 (2024).
Frenkel, M., Corban, JE, Hujoel, MLA, Morris, Z. & Raman, S. Oncofusion PROD-ATAC. GitHub https://github.com/mfrenkel16/OncofusionPRODATAC (2024).
Discover more from Tamfis Nigeria Lmited
Subscribe to get the latest posts sent to your email.



 Hot Deals
Hot Deals Shopfinish
Shopfinish Shop
Shop Appliances
Appliances Babies & Kids
Babies & Kids Best Selling
Best Selling Books
Books Consumer Electronics
Consumer Electronics Furniture
Furniture Home & Kitchen
Home & Kitchen Jewelry
Jewelry Luxury & Beauty
Luxury & Beauty Shoes
Shoes Training & Certifications
Training & Certifications Wears & Clothings
Wears & Clothings
















